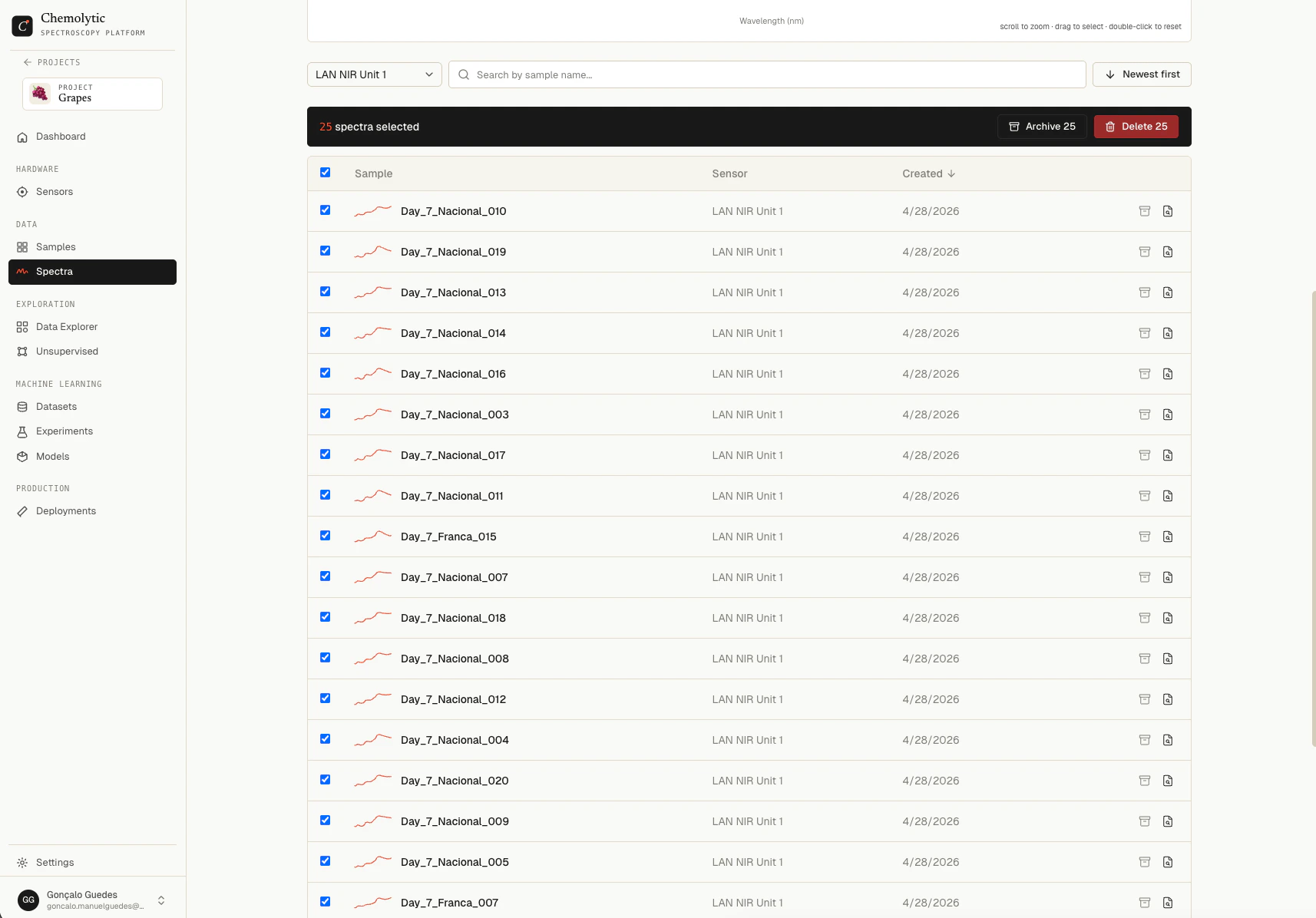
A spectrum is one measurement from your sensor: a curve of intensity values across wavelengths or wavenumbers. Spectra are the raw input to all model training in Chemolytic.Documentation Index
Fetch the complete documentation index at: https://docs.chemolytic.com/llms.txt
Use this file to discover all available pages before exploring further.
Spectra page
Go to Spectra in the project sidebar.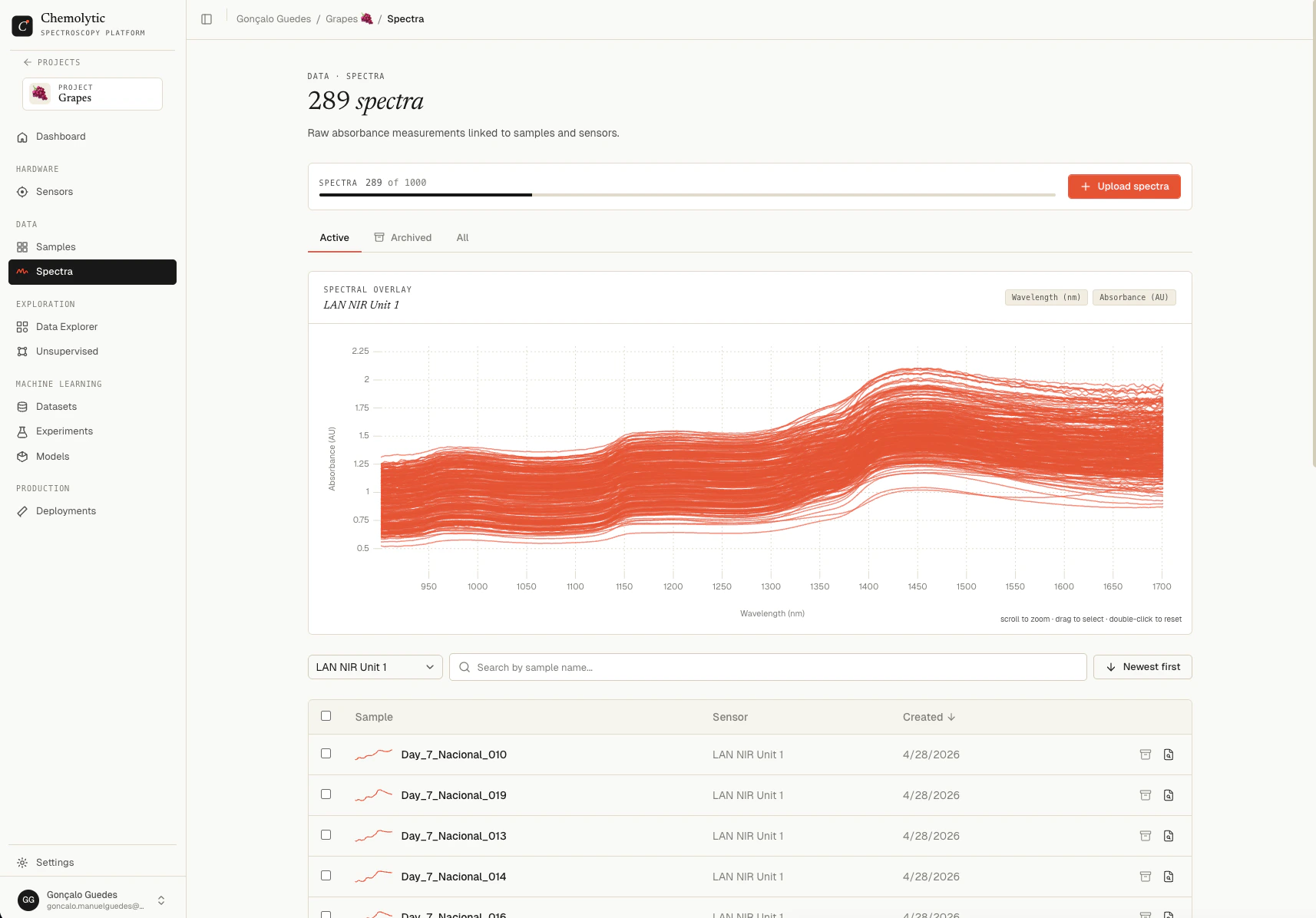
- Filters: sensor selector, search box, archive tabs (Active / Archived / All)
- Chart preview: shows all spectra for the selected sensor overlaid
- Table: one row per spectrum with sample name, sensor, creation date, and a sparkline preview
Sensor filter
The sensor dropdown is the most important filter. Each sensor has its own x-axis, so you cannot meaningfully overlay spectra from different sensors. The chart preview only appears when a sensor is selected.We’re working on multi-sensor plotting. The first version will support overlaying spectra from sensors of the same model (since they share the same x-axis). A later release will support overlaying across different models with automatic alignment.
Archive tabs
| Tab | Shows |
|---|---|
| Active | Non-archived spectra (default view) |
| Archived | Archived spectra |
| All | Both active and archived |
Search
Type in the search box to filter the visible rows by sample name.Sort
Toggle between Newest first and Oldest first using the sort button.Plan limits
The plan limit on spectra per project is shown at the top. Archived spectra count toward the limit. Delete archived spectra you no longer need to free up quota.Viewing a spectrum
Click any row in the table to open the spectrum detail panel.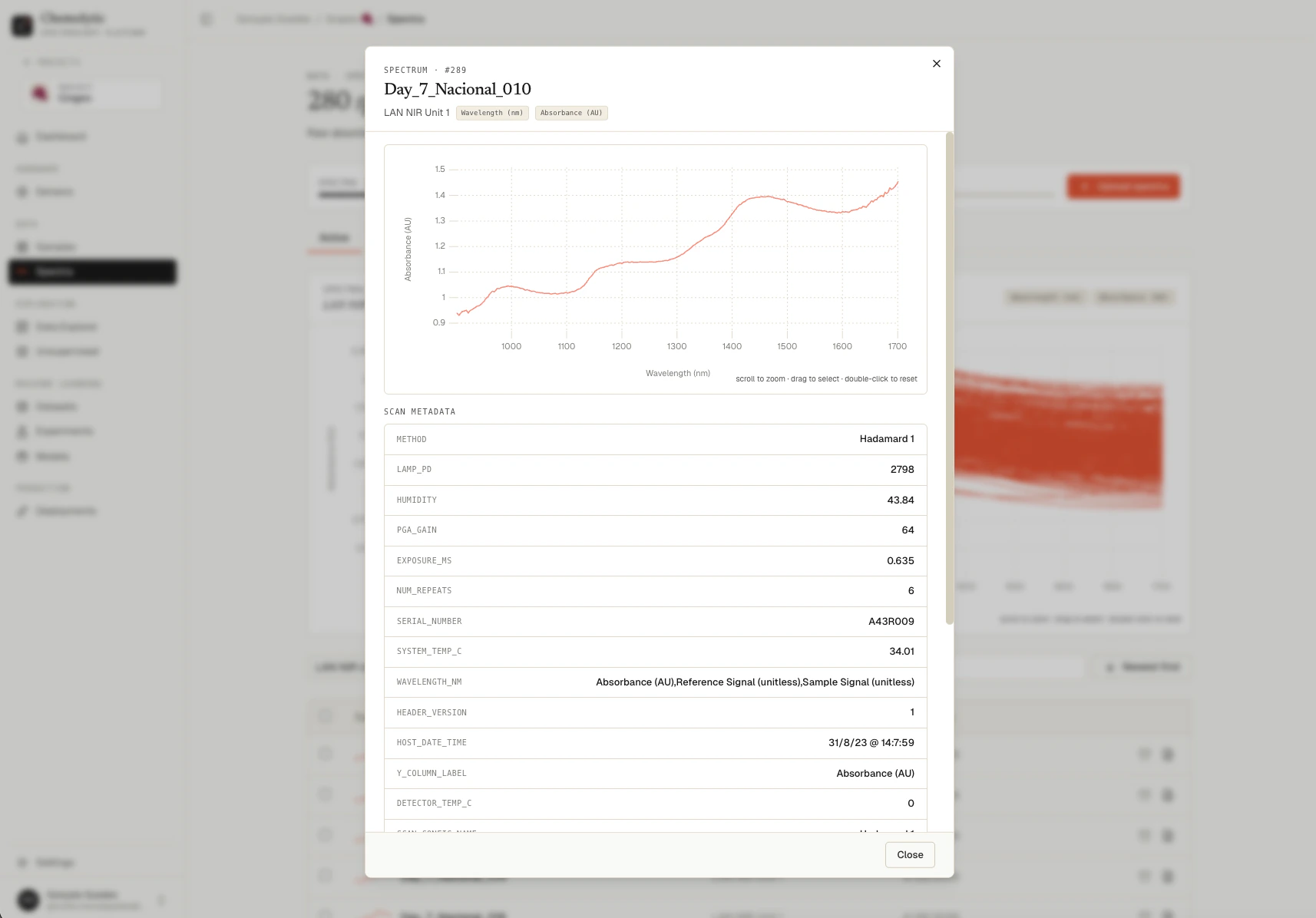
- The interactive chart for this spectrum
- Sample name and linked sensor
- Sensor units (x and y)
- Metadata extracted from the original file (if any)
- Creation date and spectrum ID
Archiving a spectrum
Archive is a reversible soft delete. Archived spectra:- Are hidden from the Active tab
- Cannot be used in new datasets or analyses
- Still count toward your plan’s spectra limit
- Can be unarchived at any time
Deleting a spectrum
Delete is permanent. Deleted spectra cannot be recovered.Bulk operations
Select multiple spectra using the row checkboxes. A toolbar appears at the bottom of the page with the count and action buttons:- Archive N (when on Active tab)
- Unarchive N (when on Archived tab)
- Delete N (when on Archived or All tab)