Once you have several runs in the same analysis, you’ll want to compare them to decide which preprocessing or method to use going forward.Documentation Index
Fetch the complete documentation index at: https://docs.chemolytic.com/llms.txt
Use this file to discover all available pages before exploring further.
Selecting runs to compare
On the analysis detail page, select multiple runs using the checkboxes in the runs list, then click Compare.
View modes
Two display modes available at the top:| Mode | When to use |
|---|---|
| Overlay | Put all runs on the same chart with different colors. Best for spotting differences quickly. |
| Side-by-side | Each run gets its own panel. Best for examining each one in detail. |
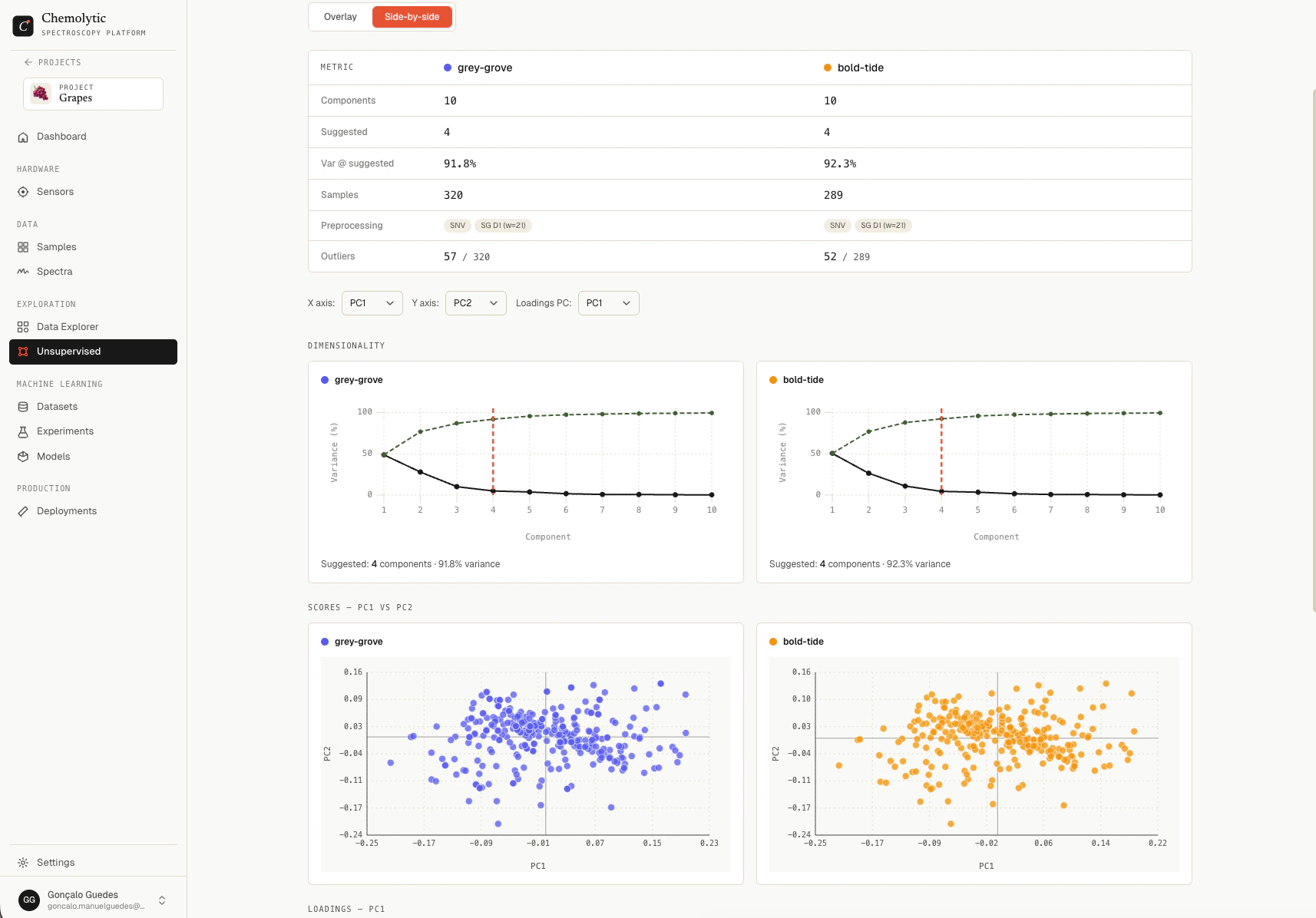
Comparing PCA runs
The compare page shows:- Metrics table: components, suggested count, variance at suggested, sample count, preprocessing, outliers
- Scree plot: explained variance per component, one line per run
- Scores plot: PC1 vs PC2 (configurable axes), all points coloured by run
- Loadings plot: spectral loadings for the selected PC, one line per run
- Outliers plot: T² vs Q residuals, all points overlaid
Comparing t-SNE runs
t-SNE compare shows each run’s 2D embedding. In overlay mode, all points appear on the same plot, coloured by run. This is rarely useful because t-SNE coordinates are not comparable between runs (different random initializations). In side-by-side mode, each run gets its own panel. This is the right way to compare t-SNE runs: look at the cluster structure of each map, ignore the absolute positions.Comparing K-Means runs
K-Means compare shows:- Metrics table: k, silhouette, inertia, sample count for each run
- Cluster sizes: bar chart per run showing how samples split